Tutorial 7: Graph Computing Benchmark
How to use this tutorial
- This tutorial is also available in Jupyter notebook format. To access and run the Jupyter notebook version of the tutorial, please sign up for free developer access by following instructions at https://github.com/juliustechco/juliusgraph.
- Additional resources (video demos & blogs) are available at http://juliustech.co.
- To report bugs or request new features, please raise an issue here. To schedule a live demo, please go to http://juliustech.co. Please check out this FAQ page or email us at info@juliustech.co for other general inquiries.
- This tutorial is copyrighted by Julius Technologies, its use is governed by the terms of use.
Introduction
Julius Graph Engine features a low-code domain specific language, RuleDSL, which makes it easy for developers to write complex business logic and also enables the efficient creation and orchestration of a computational DAG. By orchestration we mean all the runtime organization and house-keeping required so that the nodes in the DAG are executed in the right order with the correct data feeds between them.
In a typical enterprise system, the entire analytical logic and data flow can be so complex that the resulting computational DAG (directed acyclic graph) becomes extremely large. Running pricing and risk for a bank's trading portfolio is one situation where the resulting graph can grow to tens of millions of nodes. It is very difficult in practice to create and orchestrate such large DAGs, which often represents the main bottleneck when implementing a graph computing solution.
In this study, we compare the performance of DAG creation and orchestration between Julius and a few other well-known graph computing packages such as Dask, Dagger.jl and Tensorflow. Even though all the solutions support parallel execution of computational DAGs, only Julius supports parallel DAG creation. So to even the playing field, this comparison is only run off a single computer without distribution.
Benchmark Setup
The problem we used for benchmarking is to compute the sum of all the even terms of a Fibonacci-like sequence:
All $y_i$ and $s_k$ here are vector of length 10, with the initial terms $y_0$ and $y_1$ being random vectors of length 10. The benchmark problem is so designed that the resulting computational DAG is both deep ($y_n$ has very long chain of dependencies) and wide (the $s_n$ has many dependencies), in order to be representative of the wide variety of computational DAGs in the real world. The following images shows the computational DAGs for both $y_{10}$ and $s_{10}$.
| Deep DAG ($y_n$) | Deep & Wide DAG ($s_n$) |
|---|---|
 | 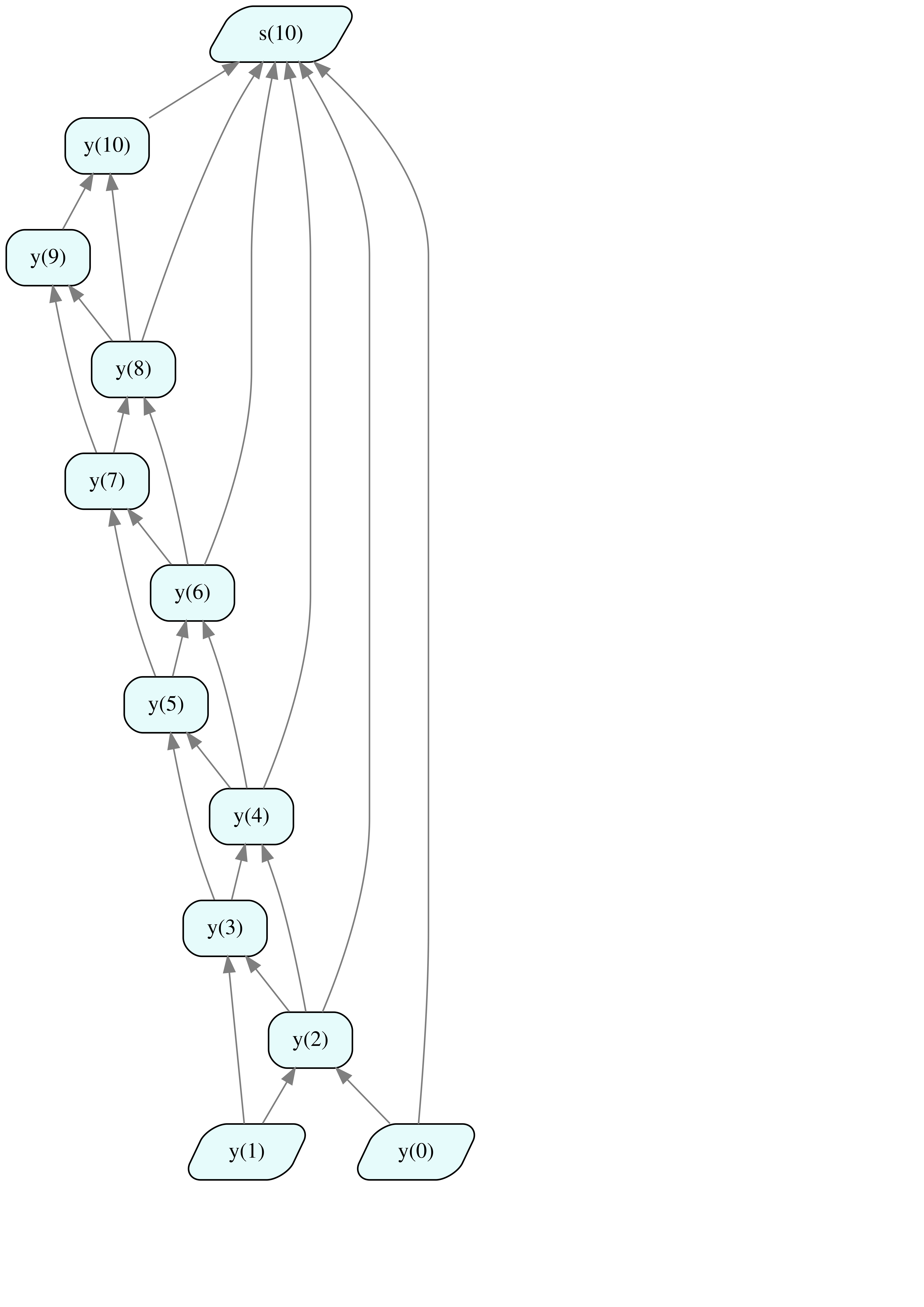 |
The methodology of the benchmarking is straightforward: we simply create and run the computational DAGs using the different graph solutions, then record their wall clock time. Python time is measured by %time, and Julia time is measured by @time. Since the numerical computation of the sequence is trivial, the time recorded is almost 100% spent on the creation of the DAGs and the orchestration of its execution.
We want to emphasize that this benchmarking study is only focused on the speed of DAG creation and orchestration. We are not testing any other features of the respective software packages. However, given that graph creation and orchestration is often the most time-consuming part of running large computational DAGs, it is of great practical interest to understand its performance characteristics.
The source code for the benchmarking is listed in the appendix.
Results
The hardware for running the benchmark is a single laptop with a 6-core intel i-7 CPU and 64 GB of memory. The following table shows the results of the benchmark for $s_n$ of different $n$, where all timing numbers are reported in seconds. The benchmark run is stopped if it did not finish under 6 hours, which is too long for practical use. Error indicate the particular run ended in exceptions before the 6 hour time limit is reached.
| $s_n$ | Dask | Dagger.jl | Tensorflow | Julius |
|---|---|---|---|---|
| 1,000 | .04 | .3 | 2.28 | 0.02 |
| 5,000 | 5.6 | 1.83 | 15.2 | 0.15 |
| 10,000 | 22 | 3.2 | 33.8 | 0.22 |
| 20,000 | 115 | 6.5 | 83 | 0.34 |
| 50,000 | 984 | 16.5 | 423 | 1.3 |
| 100,000 | error | 33 | 2,093 | 2.6 |
| 200,000 | error | error | 10,023 | 4.3 |
| 500,000 | error | error | > 6hr | 15 |
Only Julius is able to complete the $s_n$ benchmark run for $n$ = 500,000 within the 6 hour limit (it only took Julius 15 seconds). In order to understand how Dask, Dagger.jl and Tensorflow failed, we also benchmarked an easier problem of computing the $y_n$, whose computational graph is deep but not wide. The results are:
| $y_n$ | Dask | Dagger.jl | Tensorflow | Julius |
|---|---|---|---|---|
| 1,000 | .04 | .3 | 2.19 | 0.02 |
| 5,000 | 5.4 | 1.55 | 11 | 0.11 |
| 10,000 | 21 | 3.2 | 23 | 0.2 |
| 20,000 | 106 | 6.3 | 49 | 0.34 |
| 50,000 | 702 | 16.2 | 121 | 1.1 |
| 100,000 | 3,258 | 33 | 280 | 2.5 |
| 200,000 | 12,539 | error | 570 | 3.6 |
| 500,000 | >6hr | error | 1,491 | 13 |
For $y_n$, Dask and Tensorflow performed considerably better than the case of $s_n$. It suggests that a wide computational DAG poses difficulties to these solutions with big performance degradation, often more than 10 times. In contrast, Julius performed extremely well in both cases, with little difference in its overall timing.
It is also relevant to show the maximum computational DAG size that can be created and orchestrated from a single computer, assuming 6 hours is a practical time limit. The Julius results below is based on an actual run of $s_n$ for $n$ = 50MM, which took less than 5 hours. The Julius performance for $n$ = 50MM is bounded by the 64GB RAM on the laptop as heavy disk swaps occurred during the run. Julius could handle much bigger DAGs with more RAM.
| Solutions | Dask | Dagger.jl | Tensorflow | Julius |
|---|---|---|---|---|
| Max DAG Size | < 100K | < 200K | < 500K | > 50MM |
In practice, a wide computational DAG with 500K+ nodes is not uncommon, so the above result suggests that Dask, Dagger.jl and Tensorflow could run into difficulties in those scenarios. In comparison, Julius can comfortably create and execute computational DAGs of at least 50MM nodes from a single computer, regardless of its shape, making it a suitable solution for large enterprise problems.
Conclusion
This benchmark clearly demonstrates Julius' huge advantage in speed and scalability. Julius can create and orchestrate DAGs with tens of millions of nodes from a single computer. Moreover, Julius' graph construction can be easily parallelized thanks to the simple syntax of RuleDSL, extending Julius' upper limit to billions of nodes if necessary.
In comparison, Dask, Dagger.jl and Tensorflow are much slower in graph creation and orchestration; neither do they support parallel DAG creation. It is critically important for developers to consider the scalability implications of different solutions for real world problems.
Appendix: Source Code
All of the code below is directly runnable once the dependent packages are installed.
The cleanest way to implement the $y_n$ sequence is via recursion. However, Dask, Dagger.jl and Tensorflow do not yet support recursive functions, so we have to write an explicit loop in their implementations. Julius' RuleDSL does support recursive definitions, which is used in the Julius implementation.
Dask implementation
import dask
import numpy as np
@dask.delayed
def fib0(n) :
return np.random.rand(10)
@dask.delayed
def wsum(a, b) :
return (.3*a + .7*b)
%%time
# %%time is a magic command, only works in Jupyter notebook
# compute s_n
f0 = fib0(1)
f1 = fib0(2)
even = [f0]
for i in range(0, 10000) :
f2 = wsum(f0, f1)
f0, f1 = f1, f2
if (i%2 == 1) :
even.append(f2)
v = dask.delayed(sum)(even)
v.compute()
%%time
# compute y_n only
f0 = fib0(1)
f1 = fib0(2)
f2 = None # results here
for i in range(0, 10000) :
f2 = wsum(f0, f1)
f0, f1 = f1, f2
f2.compute()Dagger.jl implementation
Dagger.jl results are obtained using the revision 5f315d1 of the master branch, which is significantly faster than the current release version of Dagger (v0.14.4).
using Dagger
fibsum(a, b)=.3 .* a .+ .7 .* b
# compute s_n
f0 = Dagger.@spawn rand(10)
f1 = Dagger.@spawn rand(10)
s = 0. # result held here
@time begin
even = [f0]
for i in 1:N
f2 = Dagger.@spawn fibsum(f0, f1)
f0, f1 = f1, f2
if i%2 == 0
push!(even, f2)
end
end
even = fetch.(even)
fetch(Dagger.@spawn sum(even))
end
# compute y_n
f0 = Dagger.@spawn rand(10)
f1 = Dagger.@spawn rand(10)
f2 = 0. # results here
@time begin
for i in 1:N
f2 = Dagger.@spawn fibsum(f0, f1)
f0, f1 = f1, f2
fetch(f2)
end
end
Tensorflow implementation
Tensorflow offers heavy optimizations for processing tensors, which is the primary data type for building ML applications. However in a general enterprise use case, not all data processing and analytics can be expressed as pure tensor manipulations. The implementation below uses python range() and list instead of tensors to mimic a general enterprise use case, and to turn off certain tensor specific optimizations in Tensorflow. These tensor specific optimizations, if applied, will result in much smaller and simpler computational DAGs than the deep and wide DAGs we want to benchmark. The implementation below is therefore not the fastest Tensorflow implementation for computing the $y_n$ and $s_n$, but it is a fair implementation to compare with other solutions for a general enterprise use case.
@tf.function
def wsum(a, b) :
return a*.3 + b*.7
# have to wrap the top level call by @tf.function, otherwise
# Tensorflow does not create the computational graph
@tf.function
def sumeven(n, f0, f1) :
# python list is used instead of tf.TensorArray in order to ensure that
# Tensorflow create a single node with many dependencies
even = [f0]
# not using tf.range() because Tensorflow automatically
# optimizes tf.range() into a single loop node instead of
# creating n number of nodes in the graph, the latter is
# the case we want to benchmark
for i in range(0, n) :
f2 = wsum(f0, f1)
f0, f1 = f1, f2
if i % 2 == 1 :
even.append(f2)
# a single node with many dependencies is created for the summation
# to ensure a wide DAG is created
return tf.add_n(even)
@tf.function
def fib(n, f0, f1) :
# use python range to ensure deep graph creation
for i in range(0, n) :
f2 = wsum(f0, f1)
f0, f1 = f1, f2
return f2
%%time
# %%time is magic command, only works in Jupyter notebook
# compute s_n
n = 1000
sn = sumeven(n, np.random.rand(10), np.random.rand(10))
%%time
# %%time is magic command, only works in Jupyter notebook
# compute y_n
yn = fib(n, np.random.rand(10), np.random.rand(10))Julius implementation
To learn more about RuleDSL syntax and the concept of Atoms, please refer to the quickstart tutorial. The ApplyFn is a convenient Atom that allows arbitrary Julia functions to be used in RuleDSL, please refer to the mapreduce tutorial for more information on ApplyFn.
using GraphEngine: RuleDSL, GraphVM
using DataScience: ApplyFn
RuleDSL.@addrules seq begin
# Val(n<=1) takes value of either Val(true) or Val(false), indicating
# whether the term n is the initial two terms (n=0, n=1) of the sequence.
# Base on this, its dependency fib(n, Val(n <= 1)) is pattern
# matched to the correct rule below.
# Alias is a special Atom that simply invokes another rule.
fib(n::Int) = RuleDSL.Alias(fib(n, Val(n <= 1)))
# the -> is a lambda syntax in julia to declare anonymous functions
# this rule defines the recursion of the sequence for non-initial terms.
# ApplyFn is a generic Atom allows arbitrary julia function to be used
fib(n::Int, isinitialterm::Val{false}) =
ApplyFn[(a, b)-> (.7 .* a .+ .3 .* b)](
fib(n - 1, Val(n <= 2)), fib(n - 2, Val(n <= 3))
)
# for initial terms, simply return a random vector
# () means the anonymous function does not have any arguments
fib(n::Int, isinitialterm::Val{true}) = ApplyFn[()->rand(10)]()
# (x...) catches all input vectors
sumeven(n::Int) =
ApplyFn[(x...)->reduce(.+, x)](RuleDSL.@ref(fib(i, Val(i <= 1)) for i in 0:2:n)...)
end
n = 100000
yn = RuleDSL.@ref seq.fib(n)
sn = RuleDSL.@ref seq.sumeven(n)
config = RuleDSL.Config();
# to compute s_n
gs = GraphVM.createlocalgraph(RuleDSL.Config(), RuleDSL.GenericData());
@time GraphVM.calcfwd!(gs, Set([sn]));
# to compute y_n
gs = GraphVM.createlocalgraph(RuleDSL.Config(), RuleDSL.GenericData());
@time GraphVM.calcfwd!(gs, Set([yn]));
This page was generated using Literate.jl.